The Language of Bodies
by Serelora
This article was originally published on Medium.
Read full article on MediumWhy the way we describe patients determines whether AI helps them or quietly dismisses them.
Luis Cisneros · CEO & Co-Founder, Serelora & EvidenceMD · April 15, 2026
Math is just a way of counting things. Language is just a way of expressing things. And in medicine, the way you express a patient determines how you count them; and how you count them determines whether the algorithm sees a person or a ghost.
Hospital administrators should pay attention to this. More than 124 peer reviewed studies, published in respected journals, cited at conferences, and used to support clinical software and training, may have been built on data representing patients who never existed.
Not anonymized. Not synthetic. Not clearly labeled as such. Just wrong. Patterns inconsistent with human biology. Numbers that don’t behave the way numbers behave when real people are sick.
Gustavo Monnerat, Deputy Editor at The Lancet Americas, traced the problem back to its source and found what anyone familiar with hospital workflow could have predicted. The datasets came from unknown origins, were pooled at scale without provenance, and then fed into machine learning pipelines as if they were trustworthy. The pipelines learned, but they learned the wrong thing.
The question was never whether AI could learn from patient data. The question was always: whose patient data, collected how, structured by whom, and verified against what?
This is not a rare edge case. This is the architecture. And if you want to understand why, you need to understand the difference between two fundamentally opposed philosophies of how clinical data should live in the world.
The Top-Down Trap: Big Data as a Confidence Game
Epic Cosmos is impressive the way a warehouse is impressive. Over 300 million patients. 19 billion encounters. 2,000 hospitals. 47,000 clinics. When you hear those numbers, the temptation is to treat them the way we treat the GDP… as if scale itself is a proxy for truth.
It isn’t.
Top-down aggregation takes data that was generated inside a clinical workflow—messy, narrative, sometimes incoherent—and pools it into a lake ex post facto. De-identified. Averaged. Decontextualized. The origin story of any individual record is gone. Whether a lab value was entered by a physician who examined the patient or copy-pasted from a prior visit is indistinguishable. Whether a diagnosis code was assigned by a specialist or a billing department working three weeks after discharge is invisible.
For population-level trends, this is fine. Epidemiology doesn’t need every record to be perfect. It needs enough records to be statistically representative. But clinical AI isn’t epidemiology. It’s making decisions about this patient, today, in this room. And for that, the provenance of every data point is everything.

Rare diseases make the problem unmistakable. When the signal appears in only one out of every ten thousand patients, aggregation does not make it easier to detect. It pushes it further out of view. Coding inconsistencies, missing context, and duplicated records do not cancel out when events are that rare. They create false patterns. The algorithm still finds a signal, but not one that is real.
A Different Grammar
This is where the language problem becomes literal.
A clinical note is, by default, a prose document. It narrates. It hedges. It sometimes rambles. It carries context in syntax and tone that no off-the-shelf tokenizer was built to understand. Feed it raw into a language model and you’re feeding it a Victorian novel when it needed a balance sheet.
We built something called MOLG—the Medical Ontology Language Graph—because we believe the way you express a patient’s clinical state fundamentally determines how well a model can reason about it. MOLG is a compact, deterministic string format that rewrites clinical documentation in the native grammar of ontology — every condition linked to SNOMED CT, every medication to RxNorm, every lab to LOINC, every procedure to CPT. No brackets, no quotes, no JSON bloat.
The results from 245 complex clinical vignettes are not incremental. They’re structural.

This matters for a reason beyond engineering elegance. It is about what gets lost in translation. When a model processes a note, every ambiguous token becomes a potential hallucination, where the model fills in what it expects instead of what is actually documented. MOLG does not compress meaning. It removes the ambiguity that invites fabrication in the first place.
Counting Deviation, Not Fabricating Risk
Clean data changes what risk modeling can be. Most clinical risk scores do something quietly dishonest… they stack individual disease burden on top of population-derived mortality baselines without acknowledging that those baselines already include average disease burden. The math double-counts. The risk inflates. Nobody notices because the output looks reasonable and nobody audited the inputs.
Our Clinical Complexity Risk framework (CCR) is built to ask a different question. Not “how sick is this patient” but “how far is this patient deviating from where someone exactly like them, same age, same demographics, same baseline biology, would be expected to be right now?”


This distinction sounds technical until you think about what it means clinically. A 78-year-old with three chronic conditions who is stable and well-managed is not the same risk profile as a 78-year-old with three chronic conditions who just lost 12 pounds and stopped filling her prescriptions. Standard risk scores might give them nearly identical numbers. CCR separates them because the deviation from expected is the actual signal.
Missing data is handled by Bayesian shrinkage rather than silent nullification. Which is another way of saying we admit what we don’t know instead of pretending we do.
The Only Antidote: Data That Lives in the Room
Serelora doesn’t sit outside the EHR as an analytics dashboard you open when you have thirty minutes to interpret visualizations. It ingests data the way care actually happens—ambient scribes, lab pulls, imaging, patient messages, claims—and returns structured MOLG records and CCR scores inside the clinical decision loop. The data that trains our models is the same data the clinician used to treat the patient an hour ago.
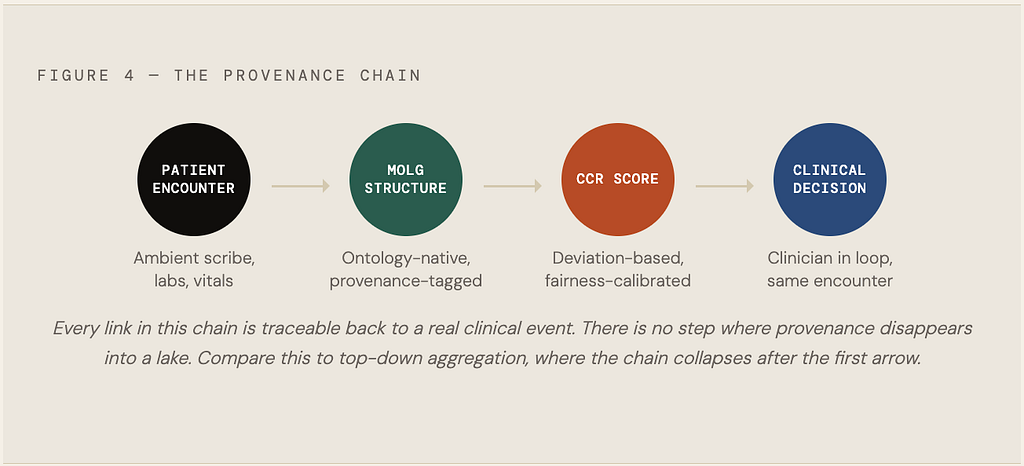
This closed loop is not a product feature. It is the only honest response to the problem Monnerat documented. If an AI model trained on questionable data is already shaping care decisions, and with more than 124 papers behind it that is a fair assumption, then the standard for trustworthy clinical AI is not more data. It is data with clear provenance, structured at the source, validated by the workflow that produced it, and explicit about its limits.
Scale without structure is not progress. It is a very large warehouse of uncertainty wearing a confidence interval as a costume.
The way we describe patients determines how we count them. How we count them determines how we filter, approximate, and predict. And how we predict—even imperfectly, even probabilistically—shapes what care they receive. That’s not a technical problem. That’s a moral one. And it starts with being honest about the language.
RELATED ARTICLES
Explore more insights and perspectives from our team.


